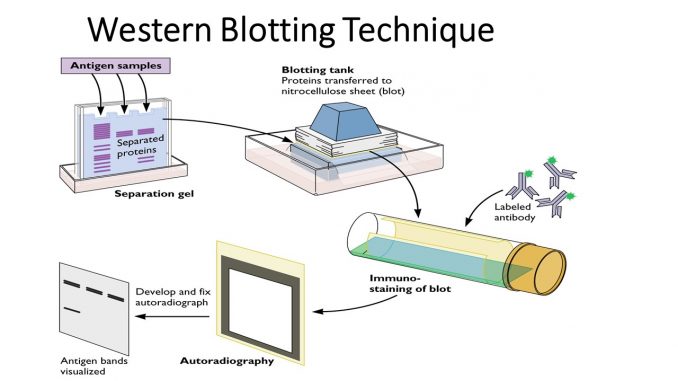
The Western blot is a widely used laboratory technique for detecting and analyzing specific proteins in complex biological samples. Its scientific basis relies on a combination of protein separation, transfer, and antibody-based detection, allowing researchers to identify proteins by their size and specific molecular characteristics. Understanding these underlying principles is essential for interpreting results accurately.
1. Protein Separation by Electrophoresis
The first fundamental principle of Western blotting is the separation of proteins using gel electrophoresis, typically SDS-PAGE (sodium dodecyl sulfate-polyacrylamide gel electrophoresis).
-
SDS and Denaturation: SDS, an anionic detergent, binds to proteins, giving them a uniform negative charge relative to their size. This denaturation ensures that proteins are separated based primarily on molecular weight, rather than shape or charge differences.
-
Polyacrylamide Gel Matrix: The gel acts as a molecular sieve, slowing the migration of larger proteins while allowing smaller proteins to move faster. This creates a size-based separation, which is fundamental for identifying immunoblotting information hub of interest.
Scientific Principle: Proteins migrate through an electric field according to their size, enabling separation for downstream detection.
2. Transfer to a Membrane
After separation, proteins are transferred from the gel to a solid membrane, such as nitrocellulose or PVDF.
-
Electrotransfer: An electric field drives proteins out of the gel and onto the membrane, where they become immobilized.
-
Protein Binding: Membranes have high protein-binding affinity, allowing the proteins to remain fixed in their separated positions.
Scientific Principle: Immobilization on a membrane creates a stable, accessible surface for antibody-based detection without disrupting the size-based separation achieved in the gel.
3. Blocking and Reducing Non-Specific Binding
Membranes are treated with a blocking solution (e.g., BSA or non-fat milk) to coat unoccupied binding sites.
-
This prevents non-specific binding of antibodies, which could otherwise produce background noise and false-positive signals.
Scientific Principle: Blocking improves specificity, ensuring that only antibodies recognizing the target protein generate detectable signal
4. Antibody-Based Detection
The core scientific principle of Western blotting is immunodetection: the use of antibodies to specifically recognize the target protein.
-
Primary Antibody: Binds specifically to the epitope of the protein of interest.
-
Secondary Antibody: Binds to the primary antibody and is conjugated to a detectable label, such as horseradish peroxidase (HRP) or a fluorescent dye.
-
Signal Amplification: The conjugated enzyme reacts with a substrate to produce a visible signal (chemiluminescent, colorimetric, or fluorescent).
Scientific Principle: The specificity of antibody-antigen interactions allows the selective detection of the target protein within a complex mixture.
5. Detection and Quantification
The final step involves visualizing the protein bands:
-
Chemiluminescence or Fluorescence: Produces light proportional to the amount of protein present.
-
Densitometry: Quantifies band intensity to measure protein abundance relative to controls or loading standards.
Scientific Principle: The intensity of the detectable signal is proportional to the amount of bound antibody, and therefore to the quantity of target protein, allowing both qualitative and quantitative analysis.
6. Key Takeaways
The Western blot is based on a combination of fundamental biochemical and biophysical principles:
-
Electrophoretic Separation – Proteins are separated by size using SDS-PAGE.
-
Protein Immobilization – Transfer to a membrane stabilizes proteins for detection.
-
Immunological Specificity – Antibodies selectively recognize target proteins.
-
Signal Amplification – Enzyme or fluorophore conjugation produces detectable signals.
-
Quantitative Analysis – Band intensity correlates with protein abundance.
These principles make Western blotting a powerful tool for studying protein expression, post-translational modifications, and molecular interactions in research and clinical diagnostics.
